|
In each case I have used this site it has provide me with a model. PHYRE2 - Protein Homology/analog Y Recognition Engine - this is my favourite site for the prediction of the 3D structure of proteins. To obtain PDB coordinates for a protein of your interest, go to the Protein Data Bank or Molecules to Go or NCBI. Peitsch (GlaxoWellcome, Switzerland) is here. With the two protein analysis sites the query protein is compared with existing protein structures as revealed through homology analysis.īackground: "Principles of Protein Structure, Comparative Protein Modelling and Visualization" by N. 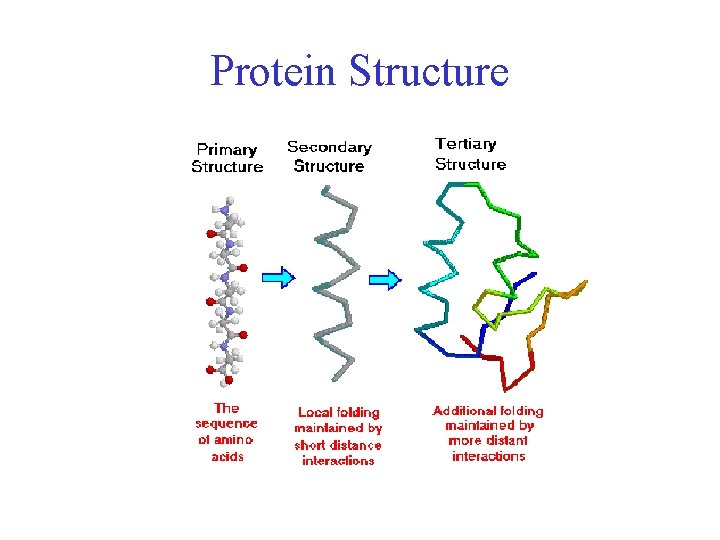
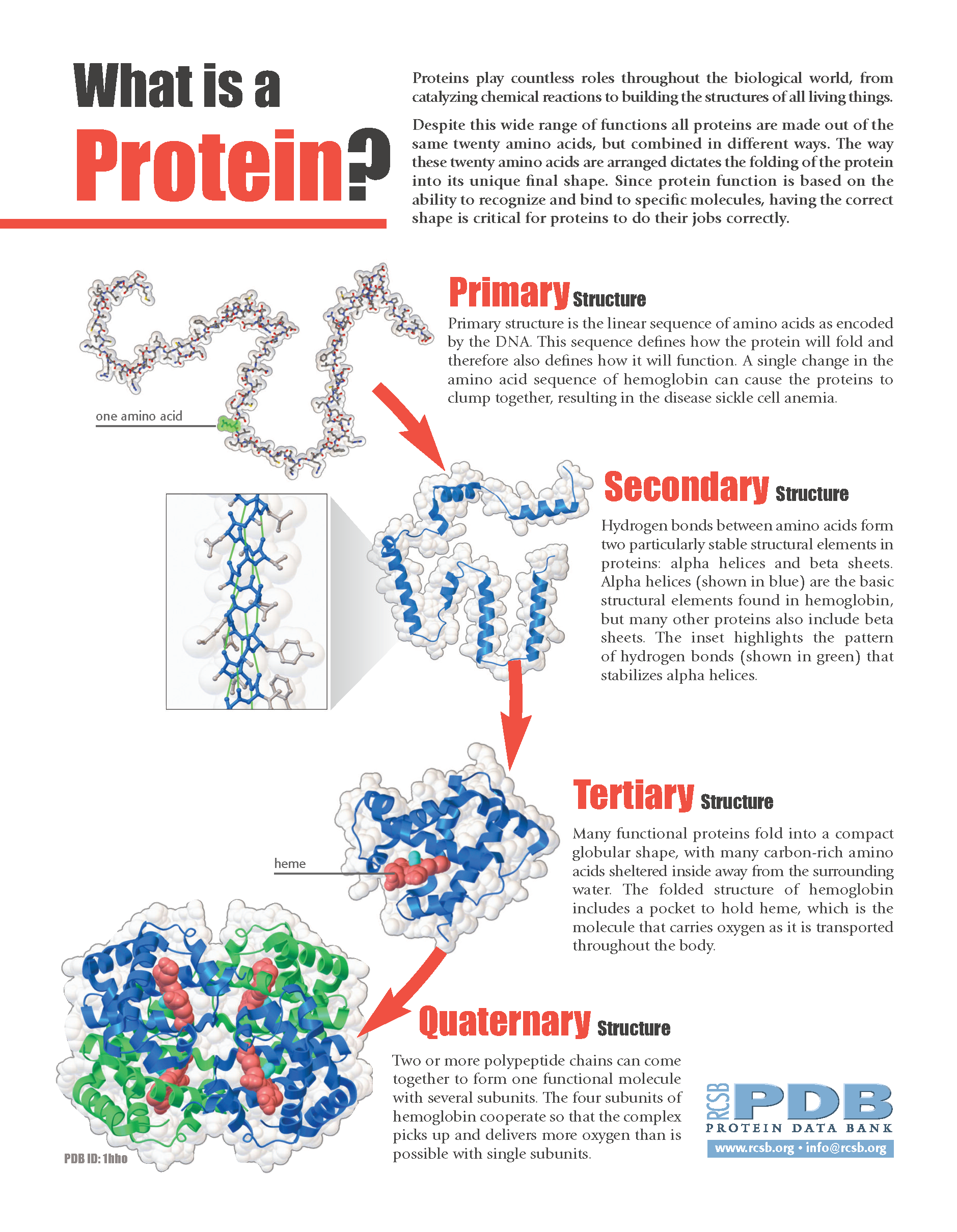
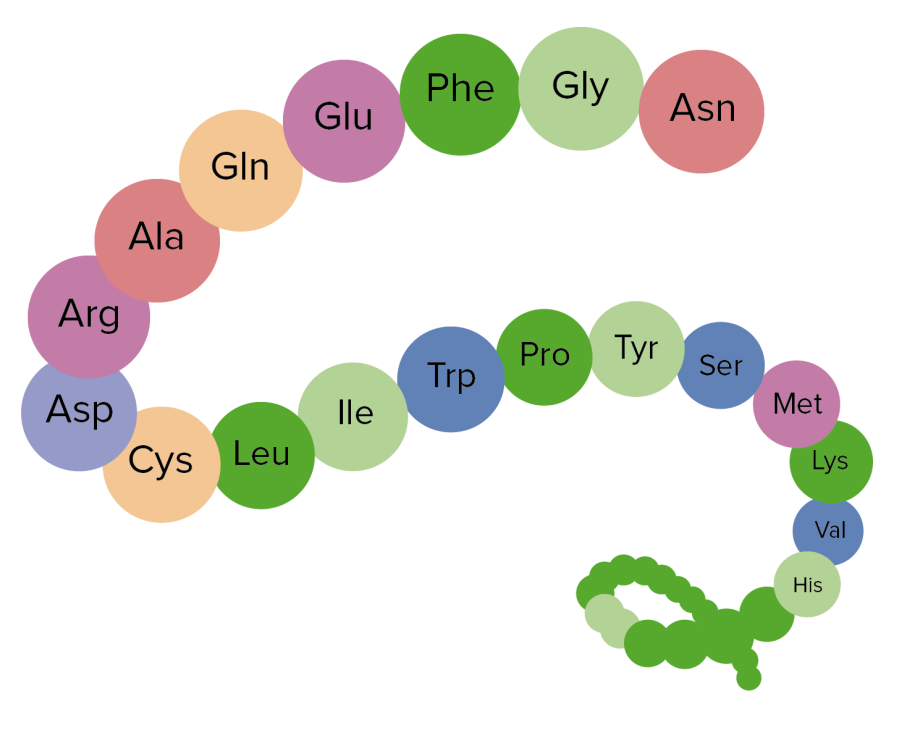
Sites are offered for calculating and displaying the 3-D structure of oligosaccharides and proteins. PDB is a standard data format that describes the protein structure.Online Analysis Tools - Protein Tertiary Structure PROTEIN TERTIARY STRUCTURE Since either the primary structure ( i.e., amino acid sequence) or the tertiary structure ( i.e., 3D folded structure) of a protein can be viewed as a graph with atom or residue as nodes and different edge construction methods.įor example, we can construct a protein from PDB file. In TorchProtein, a protein is viewed as a special case of the general graph in TorchDrug,
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
